Intro
Welcome to the 3DM walkthrough. A 3DM system is a data integration platform, we collect and integrate data about a protein superfamily.
...
If you want to use 3DM's protein visualization features you will need Yasara or PyMol and the 3DM plugin (installation instructions). This tutorial uses Yasara to demonstrate the visualization options.
System selection page
Now go again to http://3dm.bio-prodict.nl and log in. You land on a page where you can select the system you'll be working on. If you have already purchased a system(s) it will be listed here. For this tutorial we will use a nuclear receptors system, which is publicly available.
...
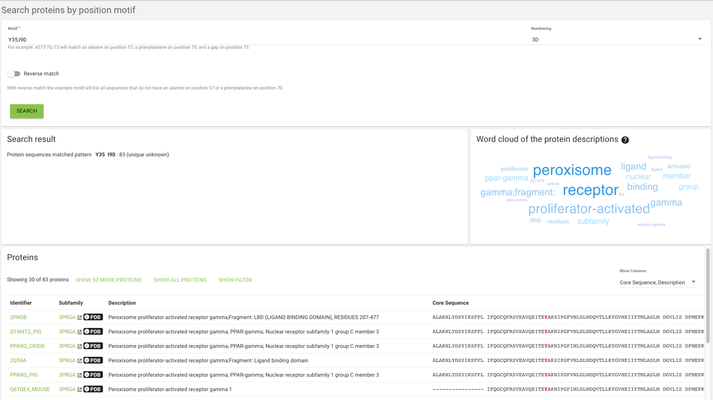
In the case of search proteins by position motif, we're looking for specific motifs on specific positions - for example we might want to find all proteins that have a tyrosine on alignment position 35 and an isoleucine on alignment position 90, then we will use the search term "Y35,I90"
Advanced
Numbering schemes
You can simplify your work with the protein of interest even more by creating a custom numbering scheme - that will cause the alignment positions to be renumbered to match the residue numbering of your protein.
...